Bioengineers recently did something incredible. During an AI search for Neanderthal proteins, they actually were able to “technically” bring some molecules back from the dead. Performing molecular de-extinction is not exactly an easy process.
To do this, the researchers applied computational methods to data about proteins from our modern humans along with our long-dead relatives/ancestors, the Neanderthals. They also included the Denisovans in this, considering the close relationship between the two others.
Doing this allowed the researchers to identify molecules that are able to kill disease-causing bacteria. Essentially, they were able to resurrect previously dead antibiotics. You can read their full research in the current issue of Cell Host & Microbe.
This is a massive discovery because it is likely we’ll be able to use this to create new antibiotics. Perhaps, we will be able to create things that can battle against infections far better than our current crop of antibiotics.
There are also some infections that standard antibiotics cannot really treat due to the bacteria being resistant to so many other antibiotics. Bringing back a long-gone version would be something this type of bacteria wouldn’t be able to defeat.
Taking A Trip To The Past

Cesar de la Fuente is the co-author of this study and a Bioengineer at the University of Pennsylvania. He had this to say about tackling current infections:
“We’re motivated by the notion of bringing back molecules from the past to address problems that we have today.”
We’ve had an issue when it comes to antibiotic development over the past few decades. While new antibiotics have been formed in that time, this has been slowing down heavily from the previous run of antibiotics that were coming out of laboratories.
Most of the antibiotics we see today have been on the market for 30+ years at this point. Yet antibiotic-resistant bacteria have been on the rise for years now, meaning a new form of treatment will be needed very soon.
Several organisms produce short protein subunits known as “peptides.” These have antimicrobial properties. A relative handful of these antimicrobial peptides are already in clinical use. Most of those have been isolated from bacteria as well.
The New…Old Antibiotic:
Of course, proteins from an extinct species could offer an untapped resource for antibiotic development. This is a realization that de la Fuente and his peers came to, funny enough, due to a movie franchise. He claimed:
“We started actually thinking about Jurassic Park.”
Rather than trying to bring back dinosaurs, however, the group asked…why not bring back molecules? That is exactly what they have done here.
To do that, they decided to do an AI search for Neanderthal proteins. Well, technically, they made an algorithm that could recognize human proteins, where they are known to be cut into peptides. To find them, the team refined the algorithm.
They applied their algorithm to available protein sequences (maps of the amino acids in proteins). Where they specifically targeted Neanderthals, Denisovans, and Homo Sapiens. The research team then used these properties of antimicrobial peptides to predict which could be new peptides that might be capable of killing bacteria.
Interestingly, finding & testing drug candidates using artificial intelligence takes just a few weeks. In contrast, it could take three to six years to do this using older methods to discover just one, single, new antibiotic.
Ancient, Long-Gone Antibiotics
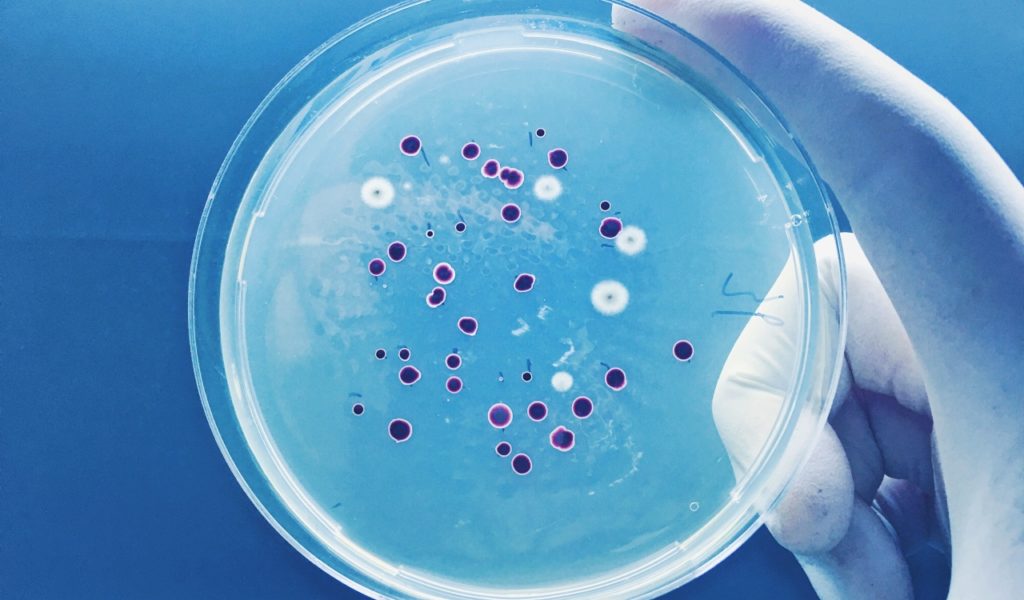
Researchers for the study tested dozens of different peptides. They checked each one to see if they could kill bacteria in laboratory dishes. When satisfied, they selected six potent peptides. Four of them are from Homo Sapiens, while one is from the Neanderthal & one is from Denisovans.
The team then gave these to mice that had been infected with Acinetobacter baumannii. This is the bacteria that has been the cause of hospital-borne infections in human beings.
All six of these bacteria halted the growth of A. baumannii which had been growing in the thigh muscle. However, none killed the bacteria. Five of the molecules killed bacteria growing in skin abscesses but it took a huge hit.
Chemical Biologist at Stanford University, Nathanael Gray, claimed that the doses used were “extremely high.” Which could have led to the issues they experienced.
Due to this, de la Fuente claimed that tweaking the most successful molecules could create more effective versions. Plus, altering algorithms could improve antimicrobial-peptide identification. Their initial AI search for Neanderthal proteins did seem to work, but refining this makes sense.
They are aiming to get fewer false positives. Plus, de la Fuente believes even though the algorithm used did not yield amazing molecules, the concept, and framework represent an entirely new avenue for thinking about drug discovery.
Meanwhile, Gray believes that “the big picture idea is interesting.”
However, until the algorithm can predict clinically relevant peptides with a much higher degree of success, Gray does not think that molecular de-extinction will have a huge impact on drug discovery.
Gray is ultimately correct here, because unless the stuff the team finds can work…what’s the point?
References:
AI in De-extinction: Reviving the Past, Shaping the Future
AI Search of Neanderthal Proteins Resurrects ‘Extinct’ Antibiotics